We’ve listed our top COVID-19 research highlights in 2020.
As Australia’s national science agency, we’ve been part of the global scientific response to the COVID-19 pandemic. Here are our top COVID-19 research moments in 2020.
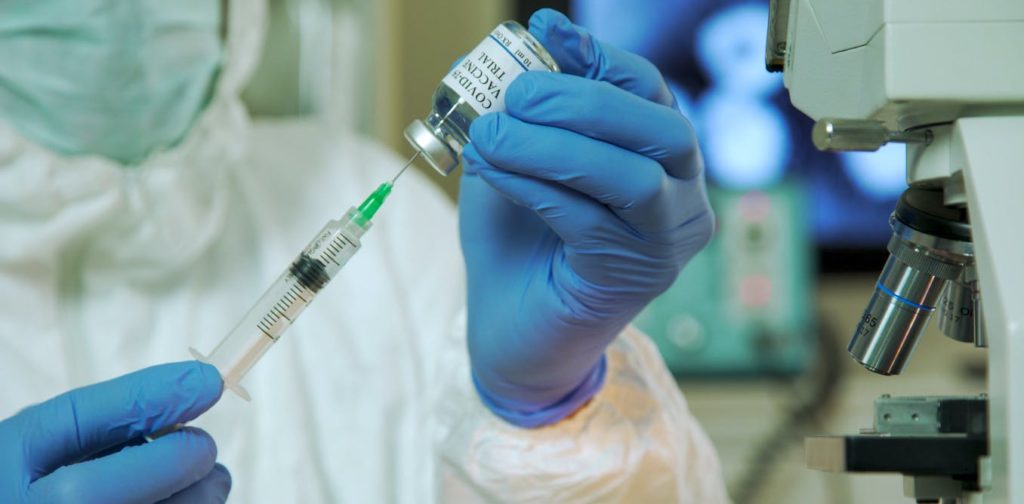
2 April 2020: We commence testing vaccine candidates at ACDP
In January, the Coalition for Epidemic Preparedness Innovations (CEPI) tasked us with studying the virus that causes COVID-19. They asked us to conduct preclinical trials for two vaccine candidates from Oxford University in the United Kingdom and Inovio Pharmaceuticals in the United States.
In April, the trials began at our Australian Centre for Disease Preparedness (ACDP). Early data from the trials has been shared with CEPI and the vaccine developers to enable them to progress on to clinical (human) trials. We’re also working with The University of Queensland (UQ) and commercial manufacturers CSL to scale-up and produce their vaccine candidate. Find out more about our role in vaccine testing.
7 April 2020: Genomics joins the fight against COVID-19
As the first step for making a vaccine we developed a novel analytics software able to capture the critical features of the virus. With this technology, we were able to compare the different versions of the virus that were rapidly emerging around the world. Like drawing a geographical map, machine learning helped us to generate an evolutionary map of the virus. By having Genomics join the fight against COVID-19, we were able to confirm that the world’s vaccine efforts were targeting the most common version of the virus.
This work was led by Dr Denis Bauer from our Health Research Program in collaboration with Dr Seshadri Vasan and the Dangerous Pathogens Team, resulting in our first COVID-19 peer-reviewed publication. Now, they have published a follow-on paper with Dr Alejandro Metke-Jimenez on methodology to link clinical outcomes to SARS-CoV-2 virus strains to enable a better understanding of COVID-19.
By partnering with GISAID, the world’s largest database for viral genomics, this technology has been scaled up to be applied to the hundreds of thousands of virus samples collected around the world. We continuously analyse the database making the resulting evolutionary landscape available as an interactive webservice for researchers to keep on top the virus’ mutational capabilities.
16 April 2020: We’re tracing sewage for early warning COVID-19 spread
Working with UQ again, we successfully found SARS-CoV2, the virus which leads to the disease COVID-19, in Australian untreated wastewater (sewage).
Led by Dr Warish Ahmed, we published a proof of concept study in April and then refined our methods and testing in June. This research showed we could identify the virus in people even before they showed symptoms. We tested seven methods to confirm the most cost-effective and rapid virus recovery process. In July, we published wastewater testing from long-haul transport like cruise ships and planes.
We continue to refine our test which provides an early warning of infection, as the virus sheds in the stools of infected people even before they show symptoms. So it’s a great early detection system for community health managers.
Have more questions? We’ve probably answered them!
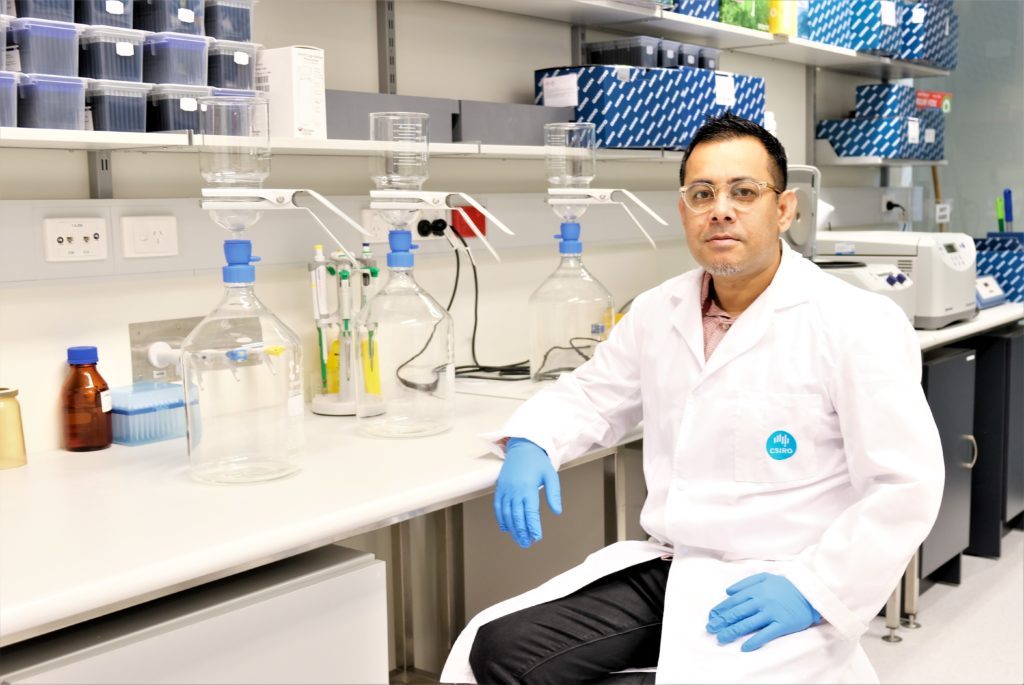

This is Dr Warish Ahmed! He researches wastewater testing can help find the presence of SARS-CoV-2 in our community.
6 August 2020: We launch Australia’s first accredited face mask testing facility
The single-use surgical face mask has become a critical piece of personal protection equipment (PPE) for our frontline workers as they fight the virus.
Although they may look like a simple apparatus, these surgical masks require significant engineering. This level of engineering is essential to ensure safe, effective and fit-for-purpose use.
In August, we launched the nation’s first accredited surgical face mask testing facility in Melbourne. The National Association of Testing Authorities (NATA) accredited the new facility.
The facility has the capacity to provide a rapid turnaround on single-use surgical face mask testing. This helps manufacturers fast-track the supply of masks for frontline healthcare workers.
Find out more about what type of masks we test and how we test them.
9 October 2020: We’re testing the G-strain of SARS-CoV-2
Most vaccines under development are being modelled on the original D-strain of the SARS-CoV-2, the virus that causes COVID-19. This was because the D-strain was the most common sequence published early in the pandemic. But now the G-strain is the most dominant strain of the virus worldwide. It accounts for about 85 per cent of published SARS-CoV-2 genomes.
There was some speculation about whether COVID-19 vaccines in development would still work against the virus.
In October, we conducted an experiment that showed vaccines developed with an older virus strain should still work against the G-strain. Our researchers took blood samples from ferrets vaccinated with the Inovio candidate (as human samples were not yet available). They tested these samples against SARS-CoV-2 virus strains, both with and without the D614G mutation.
Samples from vaccinated ferrets showed a strong immune response against the virus. They developed neutralising antibodies… even against the G-strain!
The results were clear: G-strain should not affect COVID-19 vaccines.
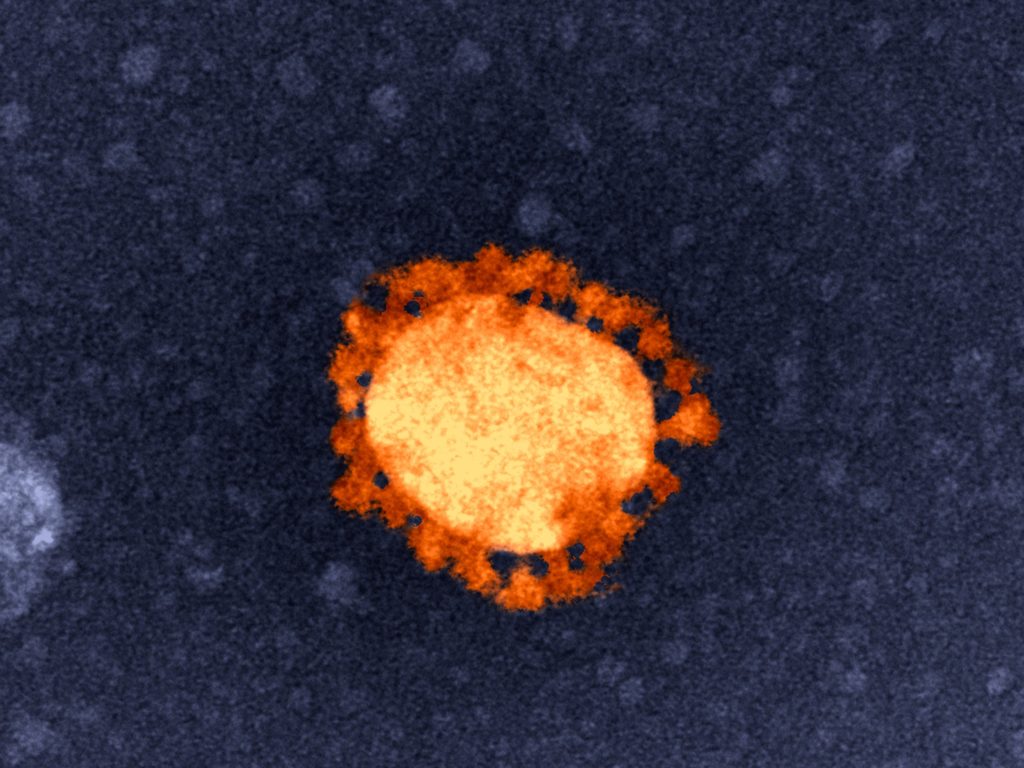
The G-strain of SARS-CoV-2 virus is the most dominant strain worldwide. But our research found the vaccines developed with the old strain should work against this new strain.
12 October 2020: We’re researching COVID-19 survivability on surfaces
In October, we published a peer-reviewed study in Virology Journal that revealed new information about the virus and how it behaves on surfaces.
We found that SARS-CoV-2, the virus responsible for COVID-19, can survive for up to 28 days on common surfaces including banknotes, glass – such as that found on mobile phone screens – and stainless steel.
To improve our understanding of how this new virus behaves, our researchers studied the survival rates of infectious SARS-CoV-2, dried in an artificial mucous solution, on six common surfaces. These surfaces were stainless steel, glass, vinyl, paper and polymer banknotes, and cotton cloth. These are examples of high contact surface areas such as glass on touchscreens and stainless steel doorknobs.
Find out more about how we conducted the study and the fascinating results.


30th November 2020 at 2:19 pm
More scientific details please n maybe less preamble?