Our researchers are studying the SARS-CoV-2 virus in order to fast track vaccine development.
As the world faces the COVID-19 pandemic, it’s critical for us to understand the genetic make-up of this virus.
We have been analysing global data on the genome sequence of the new coronavirus SARS-CoV–2 to fast track our understanding of the disease. The findings will help researchers around the world understand how strains of the virus evolve and identify new clusters of the virus.
The genome of SARS-CoV-2 is not changing as rapidly as other RNA viruses like influenza. But we need to monitor changes in the virus that can potentially modify the way it causes harm to its hosts. This is important when developing and evaluating vaccines, therapeutics and diagnostic tests.
Dr S.S. Vasan is leading our SARS-CoV-2 virus work and vaccine evaluation studies. “At this time, we do not expect it will affect the development and evaluation of COVID-19 vaccines, therapies and diagnostics. But it is important information to monitor as preclinical and clinical studies progress,” he said.
Virus decoded with genomics
Genomics, or genome sequencing, is the process of figuring out all the information encoded in the structure of an organism.
Coronaviruses are made up of RNA instead of DNA. This means they mutate and change quickly. These mutations make it hard to test which treatments might work against the virus.
However, identifying the mutations – or differences – among the 30,000 letters of the viral genome is not an easy task.
Understanding genome sequences helps researchers choose the right strain of the virus for vaccine and diagnostic efforts.
New software helps to select the most representative strain
Our team has developed software that analyses the genetic codes – or the blueprint – of the virus.
Our researcher Dr Denis Bauer said as the virus evolves, this blueprint becomes increasingly important. It holds instructions about the behaviour of the virus and what kind of disease it can cause.
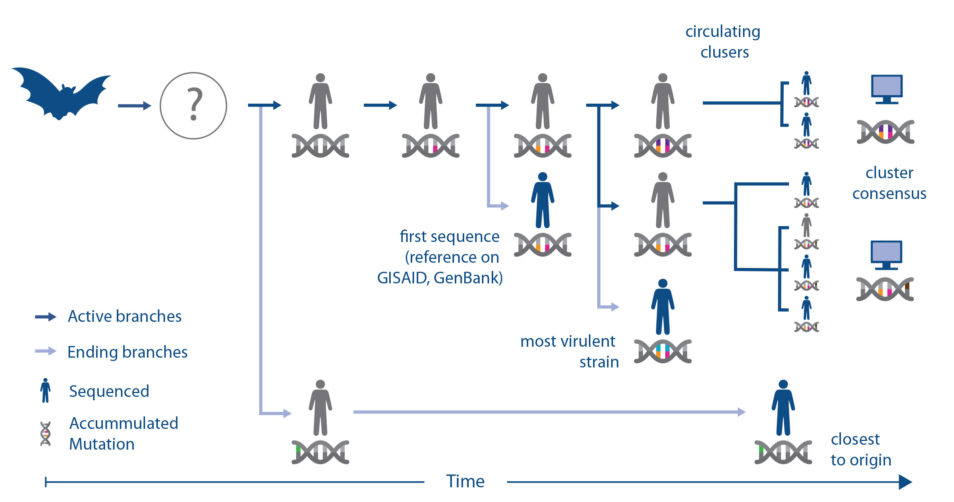
“The software developed by our team provides a visualisation showing the relationships between each strain of the virus. This helps to showcase information about the different strains of the virus. For example, where it came from and how well it might replicate,” Dr Bauer said.
It’s critical to understand the distance between the original reference point of the virus and the newly emerging components. This helped us determine which strain to choose for pre-clinical testing. The right strain for selection should have a small genetic distance to all the newly emerging components.
Analysing a significant amount of genome sequencing data in this way can help fast track our understanding of this complex disease.
Dr S.S. Vasan said after studying SARS-CoV-2’s genome sequence, we have confirmed the virus is evolving into a number of distinct clusters in different parts of the world.
“However, at this time we do not think it will affect the development and evaluation of COVID-19 vaccines, therapies and diagnostics. But we will continue to closely monitor the situation,” Dr Vasan said.
Help us better understand the COVID-19 genetic code
This work is just the tip of the information iceberg in understanding the COVID-19 virus. We are calling on the international research community to share de-identified clinical information about the disease progression alongside the genomic sequences of the virus. We need to be well informed about the genetic differences of the virus. This way we can understand the likely consequences on the progression of the disease. Then we can better tackle the disease with diagnostics and treatments.
The peer-reviewed research paper, Supporting pandemic response using genomics and bioinformatics: a case study on the emergent SARS-CoV-2 outbreak, was accepted for publication by the Transboundary and Emerging Diseases journal on 7 April 2020.


27th May 2020 at 9:46 pm
Vaccines and treatment alternatives are extreme money makers and it is likely that those who sit on key information will not share it. I hope that there will be a solution that enables us all – globally – to benefit from the research,, and that this could be done on a worldwide basis, focusing more on all humans than on the profit. The article seems to be written in this vein and I wish that all other researchers and governments too will think along these lines. I wish you – and thereby us – huge success!
27th May 2020 at 2:24 pm
The development of treatments and vaccines for emerging pathogens needs to be funded by governments, not companies as companies will only research guaranteed money makers. There was a corona virus vaccine being developed some 10 years ago but it was dumped because “there was no urgency”. What do we do if we get an Ebola like pathogen that is airborne. Currently there are no reliable treatments because it’s an Africa problem. Only by doing the work when there is no urgency can we be assured of having a solution when it’s needed.
24th April 2020 at 5:00 pm
Thank you for the incredible research you are doing and hopefully the Government will start to reverse some of extreme cuts that have happened to your budget over the years