Researchers sleuthed the human genome to find the genes most vital for viral replication. They found blocking the fibrillarin region stemmed the replication of many virulent types of viruses.
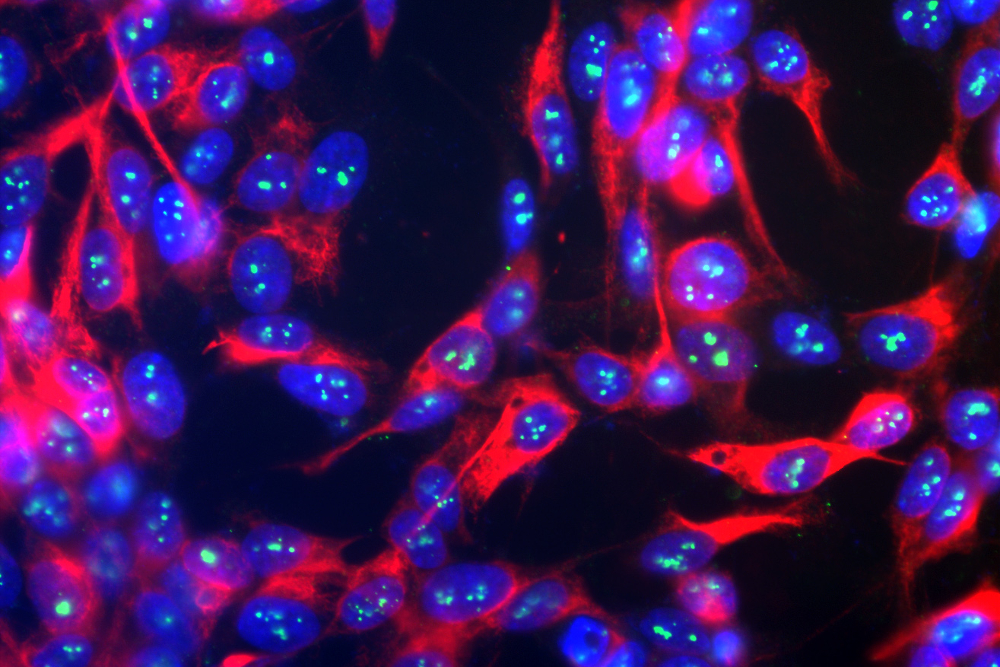
Fibrillarin stain of human neuroblastoma Feature ration
Neuroblastomas, wherein the fibrillarin enzyme is stained green. Wikimedia Commons/Gerry Shaw.
Imagine actually trying to find a needle in a haystack. Well that’s what we set out to do in our world first study to identify which proteins were responsible for helping some of the deadliest viruses — like Hendra virus— replicate and then infect people.
You know, viruses are pretty rude if you think about it. Not only are they a major source of disease, pestilence and your run-of-the-mill global misery, but what’s worse, they actually turn us into vehicles for their nasty deeds. Many viruses, including some of the most deadly, have a very limited genetic capacity. For instance the Hendra virus genome encodes only 9 proteins. Compare that to the human genome which encodes over 20,000 proteins. It’s a big difference.
With so few genes, most viruses cannot replicate unless they’re inside a host, wherein they hijack and exploit key cellular elements to multiply. If we were to identify the host proteins each virus required for infection, we’d have around 20,000 to test — a mammoth undertaking. It requires going through the entire human genome, silencing each protein-encoding gene one by one, and observing whether the virus can grow in cells lacking that particular protein. If we learned which host proteins each virus needed for infection, we’d not only further our understanding of virus biology but also identify targets for antiviral therapeutics for diseases where there are currently no treatments.
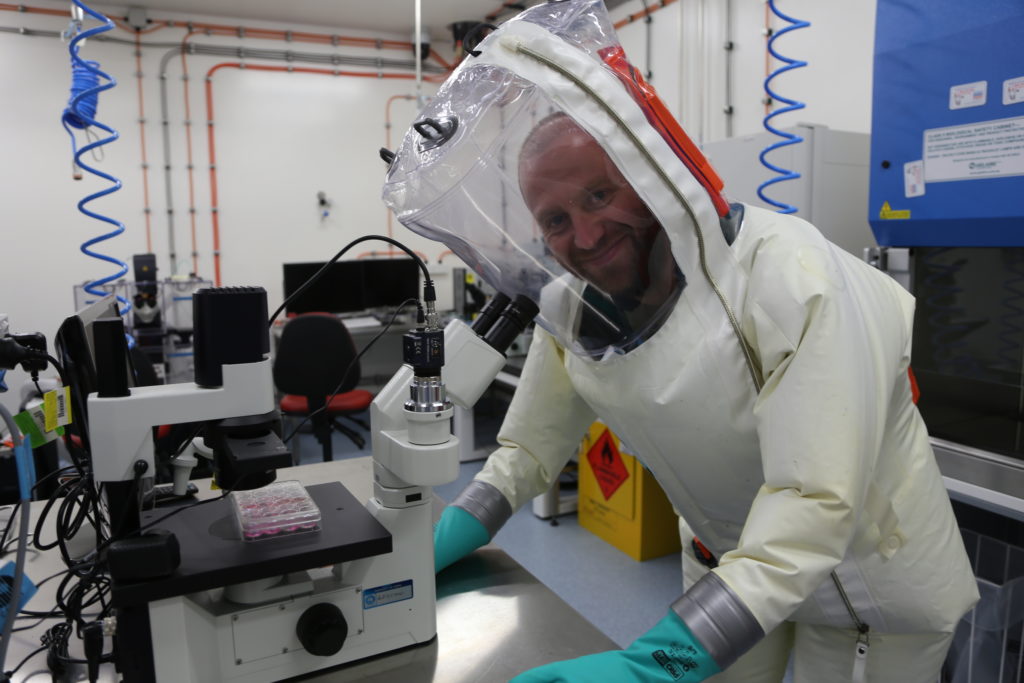

Dr Cameron Stewart looking over samples.
In response to the large number of Hendra virus outbreaks in Australia in 2011, a team at our Australian Animal Health Laboratory (AAHL) teamed up with the Victorian Centre for Functional Genomics to identify host proteins required for Hendra virus infection. Not only was this effort the first screen of its kind for a biosafety level 4 pathogen (requiring the highest level of biocontainment), it was also the first screen performed for a paramyxovirus. Some of Hendra virus’s cousins include measles, mumps, respiratory syncytial virus and human parainfluenza viruses, which collectively infect hundreds of thousands of people each year. The large collaborative study also involved researchers from CSIRO Manufacturing, Duke NUS (Singapore), the University of Georgia and Centres for Disease Control and Prevention (United States).

Researchers involved in finding Fibrillarin
The team at the AAHL finding fibrillarin.
After years of hard work, the completed screen revealed that Hendra virus infection relies on over 500 host proteins. Of all the molecules tested, the largest impact was observed upon targeting the protein fibrillarin, an enzyme that resides in the host cell nucleus. Blocking fibrillarin decreases Hendra virus infection by over 1000-fold, to the extent where virus growth is barely detectable in cells lacking fibrillarin. Closer examination revealed that fibrillarin (which resides in the host cell nucleus) is essential for the synthesis of viral RNA and viral proteins, events that occur in the host cell cytoplasm. This surprising result suggests that host proteins within the nucleus play unappreciated roles in viral replication, and may explain why so many viruses encode proteins that access the host nucleus during infection.
Finally, it was found that blocking fibrillarin impairs infection by a wide range of paramyxoviruses, suggesting that targeting fibrillarin therapeutically may be a novel avenue to blocking virus infection. Our current work is focussed on understanding exactly how fibrillarin facilitates virus replication, expanding our virology work to assess whether fibrillarin is required for other deadly viruses, including Ebola and Rabies viruses, and developing the world’s first small molecule inhibitors of fibrillarin function. Ultimately our goal is to develop a broad-spectrum inhibitor of some of the world’s most deadly viruses.


2nd July 2018 at 7:20 am
Dr Stewart, the fibrillarin enzyme should be name the Charon protein as it carries the dead to an afterlife.
2nd July 2018 at 7:16 am
The contest to name the gene has only one possible answer: the Charon Gene, it ferries the dead to an afterlife to live on.
18th May 2016 at 10:11 pm
Absolutely amazing science; a vanguard that can prevent catastrophic epidemics which could cause untold harm to animal life forms. I cannot imagine how virologists manage with such miniscule subjects.