A crystal image of a breakdown pathway of atrazine
The International Year of Crystallography is drawing to a close, and we’re not going to let it finish without showing you something about what crystallographers do. Which is not what most people would assume when they hear the word: there are crystals involved, but it’s not exactly the study of crystals as we generally think of them. It’s the study of how matter is organised, using crystals as a tool.
Now, naturally we want to know how matter is arranged. Apart from being very, very interesting to find out about, it also helps in many different fields, from drug delivery to materials science. In fact, it was crystallography that provided – controversially – the key to understanding the structure of DNA.
So assume you want to look at something in the greatest possible detail, seeing its smallest possible components. Obviously, you’d use a microscope. But there’s a limit to the smallness of things you can see that way: the wavelength of the light human eyes see. Visible light has a frequency of between roughly 400 and 700 nanometres, and can’t detect atoms, which are separated by 0.1 nanometres. This is the perfect frequency for X-rays.
We can’t make appropriate X-ray lenses to make x-ray microscopes to study molecules: we have to do it in a roundabout way. We beam X-rays onto crystals, scattering the rays, in just the same way that light reflects when it hits an object. Then we use a computer to reassemble the rays —the diffraction pattern —into an image. The diffraction of a single molecule would be so weak that we couldn’t get any meaningful information from it, so we use crystals, which have many molecules in an ordered array, to amplify the signal so we can see it. Crystals are highly ordered structures, made up of 1012 or more molecules, makes the x-ray diffraction patterns — the main tool of crystallography —possible to analyse.
Crystallographers were among the first scientists to use computers, and used them to do the advanced calculations needed to reassemble diffraction patterns into coherent images. That’s why it seemed fitting to name our supercomputer after the founders of crystallography – Lawrence and Henry Bragg. Lawrence was the first person to solve a molecular structure using x-ray diffraction.
Today we can not only view molecules in 3D, but also study the way they operate. Improvements in x-ray machines have also led to synchrotron facilities, which can produce far more efficient and precise beams.
And speaking of synchrotrons …
One of our crystallographers, Tom Peat, has deposited more than 120 structures in the Protein Data Bank using data collected at the Australian Synchrotron. They were all derived from crystals developed in CSIRO’s Collaborative Crystallisation Centre.
This is one of our favourite structures.
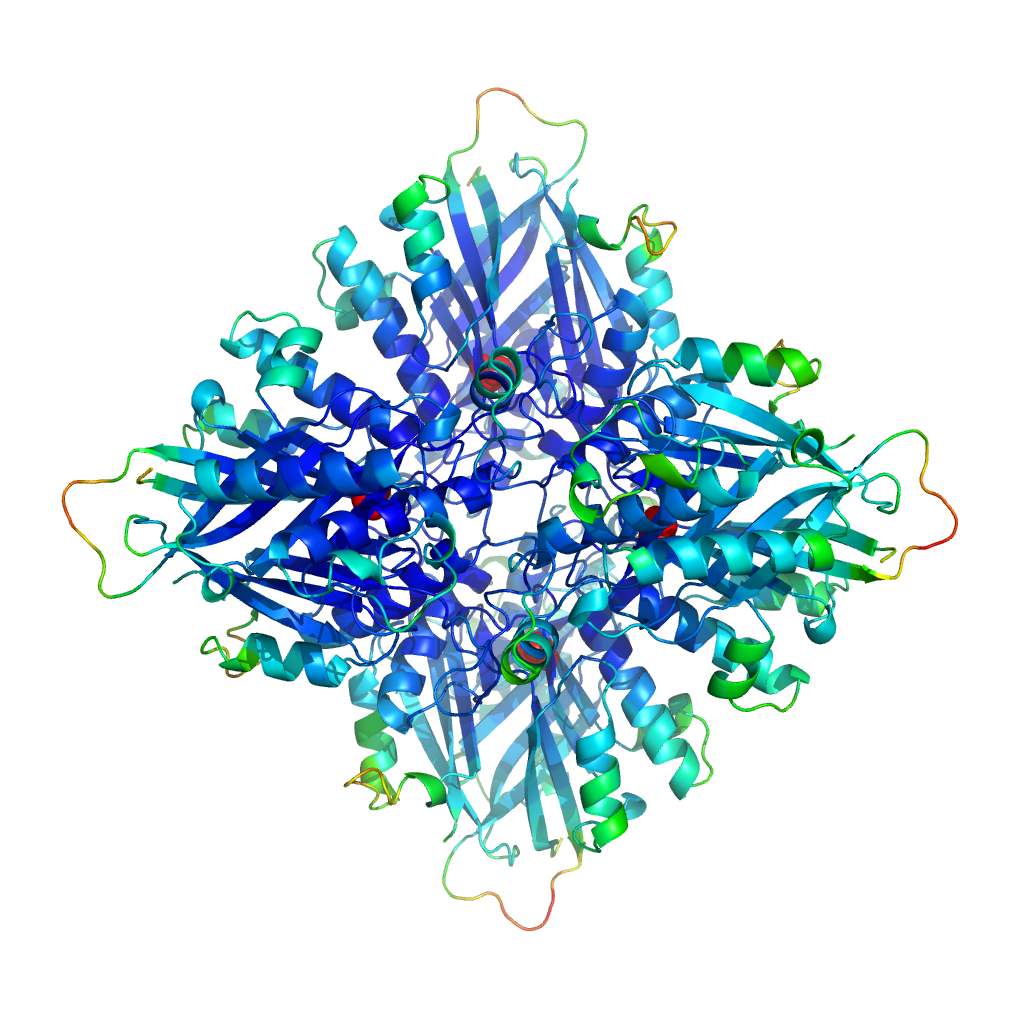
AtzF structure (the Australia crystal)
It’s the structure of AtzF. This enzyme forms part of the breakdown pathway for atrazine, a commonly used herbicide. We’re trying to understand enzymes better and use them for bioremediation – cleaning up environmental detritus such as pesticides and herbicides – and we’ve now solved the structures of four of the six enzymes involved in the atrazine breakdown pathway. We also look at protein engineering, to see if we can make these enzymes even more effective at cleaning up the environment.
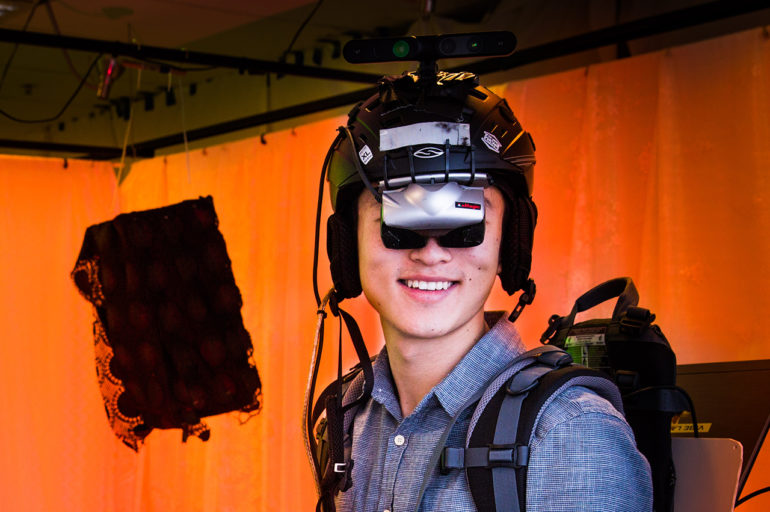
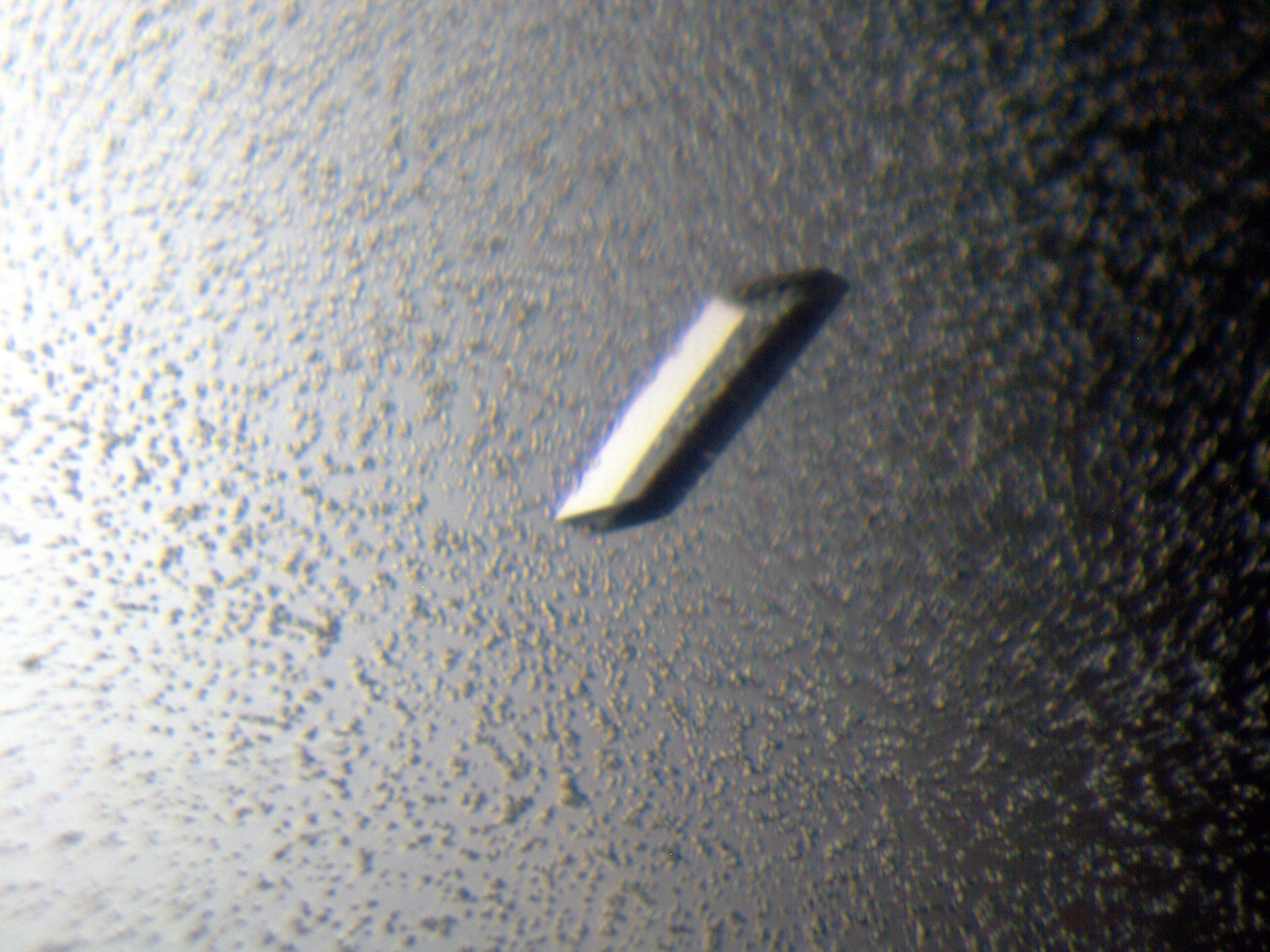
Before we get to the crystal image, there are other steps on the way. First, someone has to grow the crystals (clone the protein, express it, purify it and crystallise it). Then it’s off to the Synchrotron to get a data set (many diffraction images in sequence). Here you can see an actual protein crystal.
An actual protein crystal
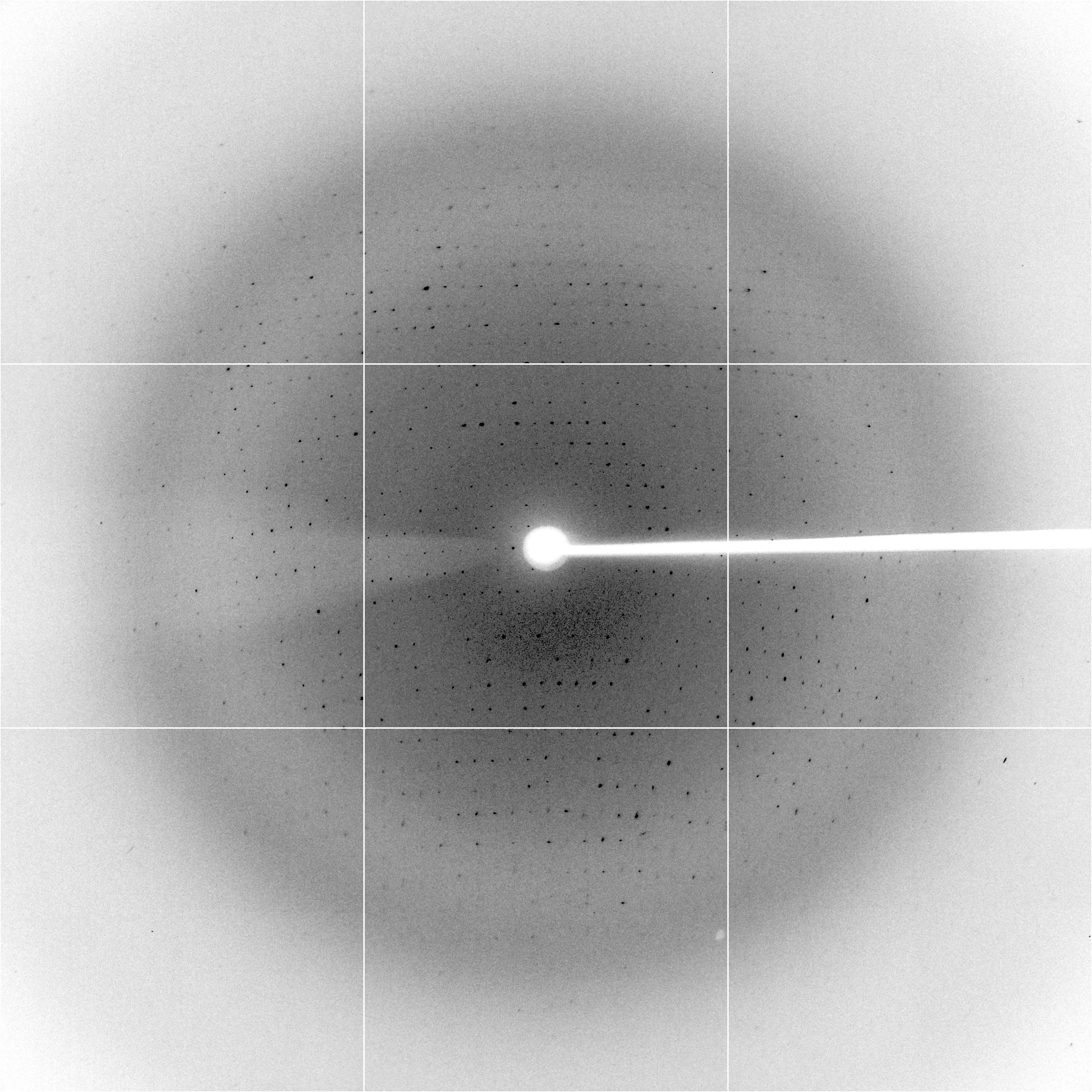
The picture on the right is the diffraction image.
Diffraction image
The crystallographers measure the intensity of the reflections (the dark dots). They combine that with the geometry and use some complicated maths (a Fourier Transform) to produce an electron density map. They then use that map to build a model.
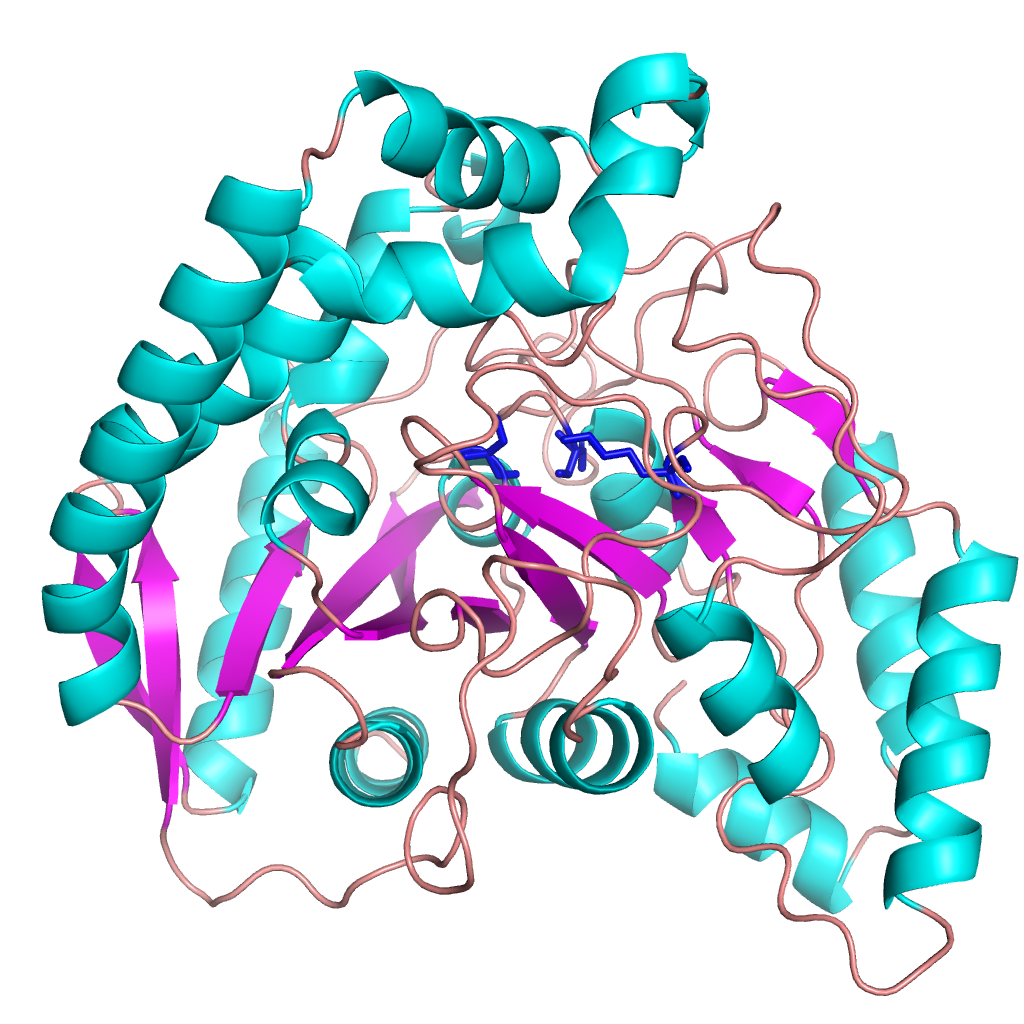
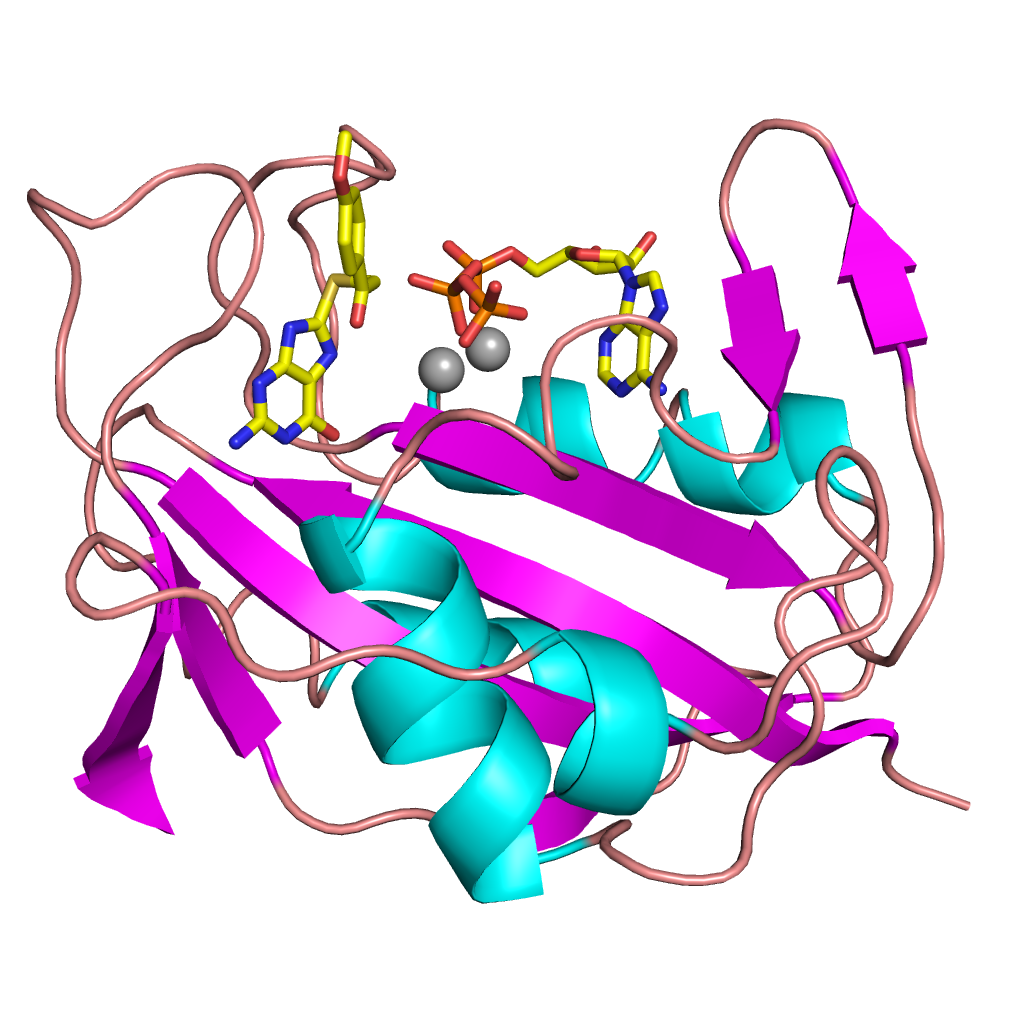
Not all our crystallography work is in the same area. We also work on some pharmaceutical applications. One of our projects, with hugely important implications for human health, is on the design of desperately needed new antibiotics. We’ve been collaborating with Monash University, looking at the pathway that sulpha drugs (such as sulfamethoxazole)– the ones we used to treat bacterial infections prior to the discovery of penicillin – take to treat Golden Staph infections in humans. The aim is to design new antibiotics that target the same pathway. You can read a paper that describes our recent findings in the Journal of Medicinal Chemistry, and here’s a picture of what we’ve been doing.
Golden Staph treatment pathway
We think this deserves its own Year. And we hope it’s clear just how important it is. Crystal clear.